Detailed instructions how to use the database.
- Browsing the database
- Searching the database
- Exporting data into CSV
- Following new records using RSS
- Adding new records
- Managing taxonomy and editing categories
- Importing multiple records
- Database feedback, requesting permissions and contributing
Browsing the database
In the top menu navigate to BROWSE, and browse the database records according to endemism (i.e. if the species is endemic to GCFR or not), life form (i.e. all records of particular life form), red list category (i.e. records of species of particular conservation status), taxonomy (displayed in tree-like form from families below), or vegetation type (i.e. records of species growing in particular vegetation type).
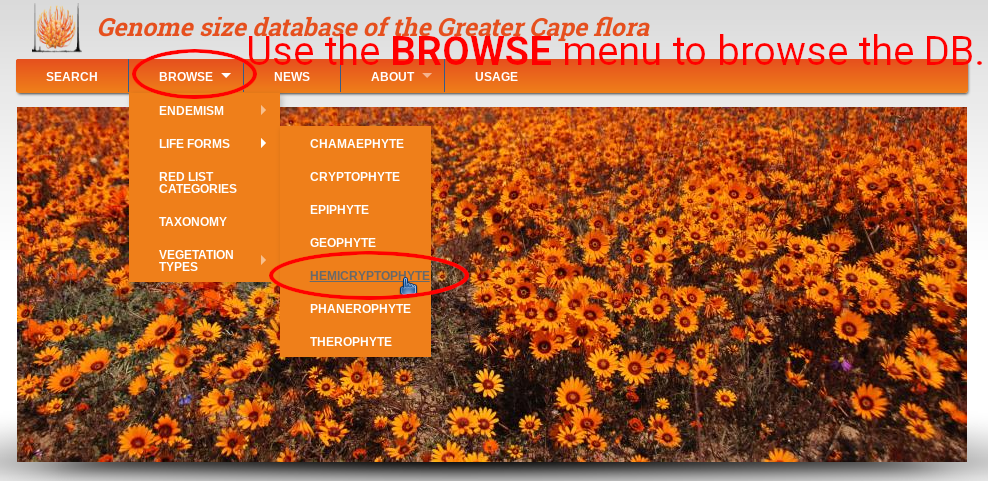
Every category contains brief description of particular vegetation type, life form, etc. Below is table with respective records, and map showing their distribution. Click to the label in the ID column to get details of the selected records. Table can be sorted by clicking on the orange column headers. Clicking to links in any other column than ID leads to table listing respective taxonomy (species), life form, or vegetation type.
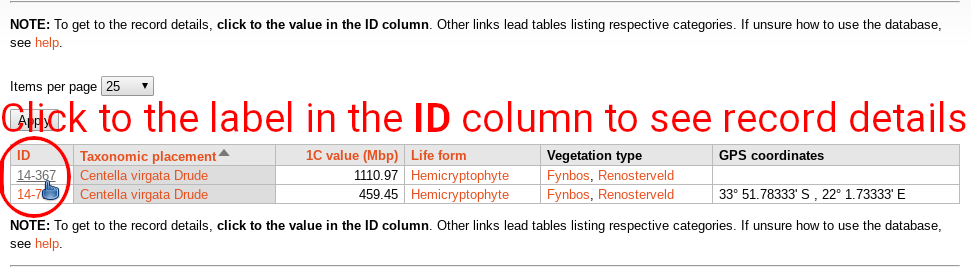
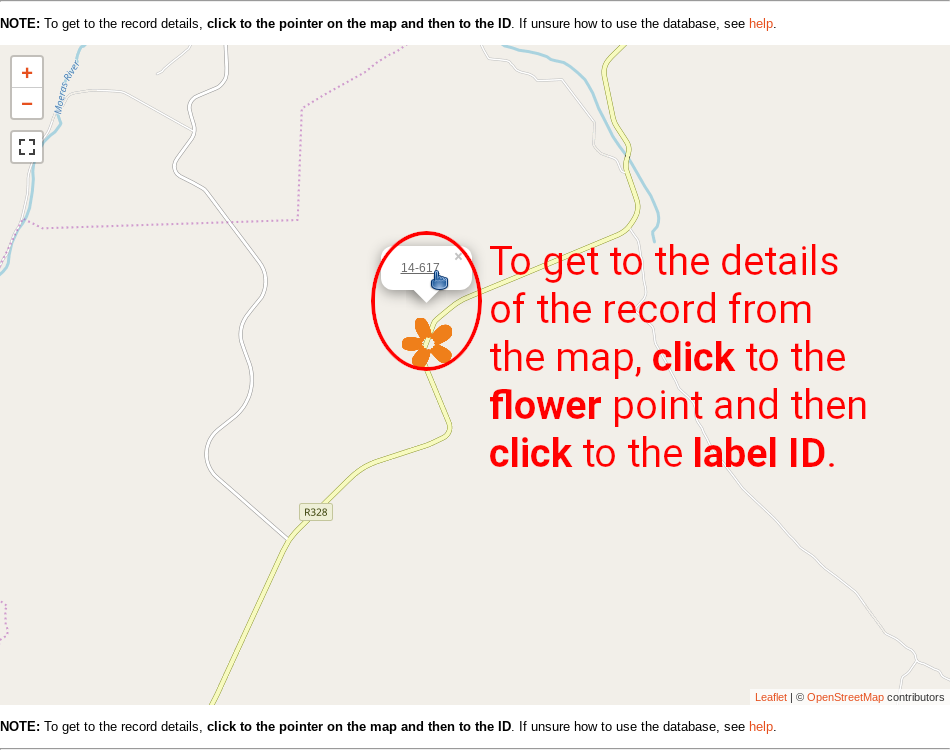
Searching the database
Basic search uses taxonomy (any rank up to family). Start to type into the search field, wait a second, and the list of matching names will appear.
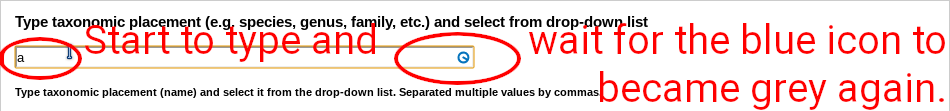
Select the taxon from the list. If there is no drop-down list, your text does not match any taxon available in the database. You must always select from the list! Otherwise the taxon search will not work.

Multiple values can be separated by commas. Start typing, select first taxon, separate any other taxon by comma, repeat.

Advanced search allows combination of various information. When using Advanced search, it’s not necessary to type any taxonomic placement.
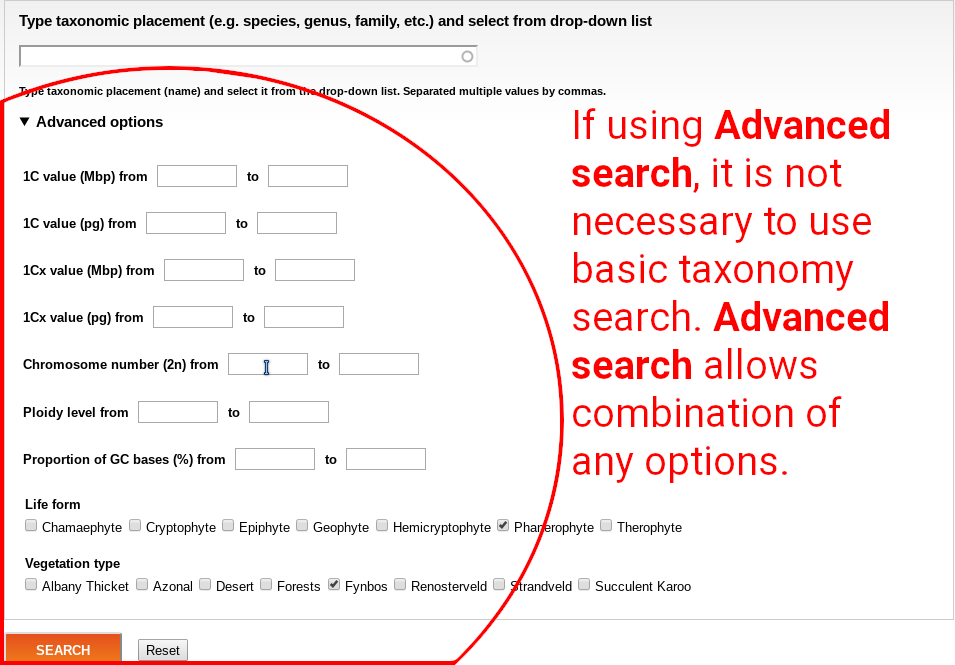
Any search still outputs same table. It is possible to go to details of any record by clicking to the label in the ID column, or export all outputs using the CSV option.
Exporting data into CSV
Any table showing browse or search results can be exported using the CSV icon. The data are exported as TSV. The export contains all data colums (not only information visible in the displayed table). The resulting text file uses UTF-8 encoding, UNIX end of lines, TAB as column separator and no cell quoting.

Following new records using RSS
New records in any category available through the BROWSE menu (see above) can be followed using RSS web feeds. The RSS icon is below the table of records.

Adding new records
X
Managing taxonomy and editing categories
X
Importing multiple records
X
Database feedback, requesting permissions and contributing
X
